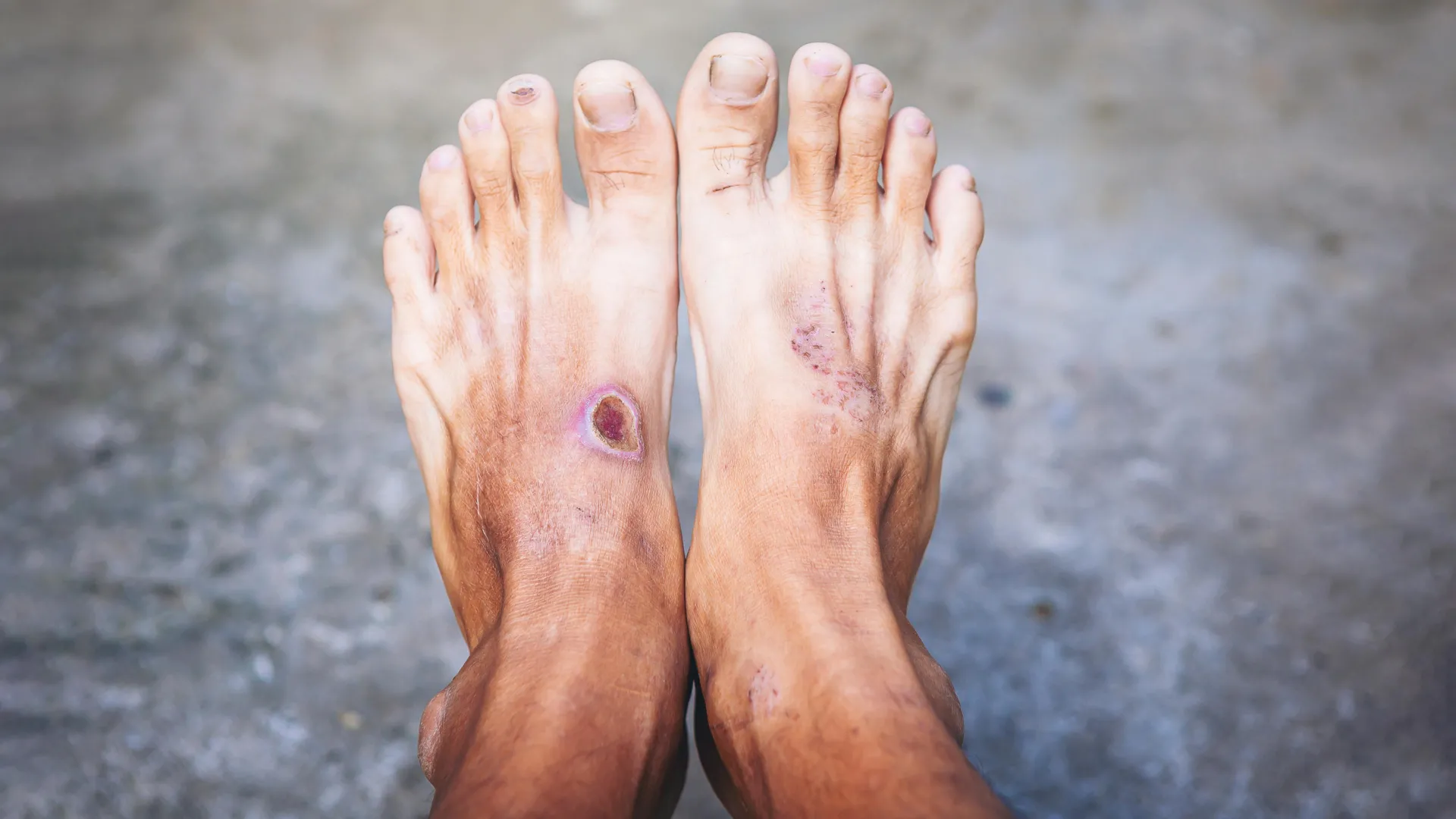
The Evolving Threat of ‘Superbug’ E. coli in Diabetic Foot Infections: A Genomic Revolution in Treatment
Over 37 million Americans live with diabetes, and a staggering 15% will develop a foot ulcer during their lifetime. But a silent, and increasingly complex, threat lurks within these wounds: E. coli. Recent global genomic research, spearheaded by King’s College London, isn’t just confirming the presence of this common bacterium, it’s revealing a hidden diversity – a vast, previously underestimated range of strains – that’s driving treatment failure and escalating the risk of amputation. This isn’t simply about a bacterial infection; it’s about a rapidly evolving genomic landscape demanding a radical rethink of how we approach diabetic foot care.
The Hidden Diversity of E. coli: Beyond the Textbook
For decades, E. coli has been recognized as a potential opportunistic pathogen in diabetic foot infections (DFIs). However, the assumption was often a relatively limited range of virulent strains. The new research shatters that notion. By analyzing genomic data from DFI samples collected worldwide, scientists have uncovered a remarkable genetic diversity within E. coli populations. This diversity isn’t random; it’s actively evolving, driven by factors like antibiotic use and the unique microenvironment of chronic wounds.
Why Does This Diversity Matter?
The answer lies in the concept of antimicrobial resistance (AMR) and virulence factors. Different E. coli strains possess varying levels of resistance to common antibiotics. More importantly, they carry different genes encoding virulence factors – the tools bacteria use to invade tissues, evade the immune system, and cause damage. A ‘one-size-fits-all’ antibiotic approach, therefore, is increasingly likely to fail against this diverse population. The study highlights that some strains are particularly adept at forming biofilms, complex communities of bacteria that are notoriously difficult to eradicate.
Personalized Medicine: The Future of DFI Treatment
The implications of this research are profound. The era of relying on broad-spectrum antibiotics for DFIs is drawing to a close. The future lies in personalized medicine – tailoring treatment strategies based on the specific E. coli strains infecting each patient. This requires rapid and accurate genomic sequencing of wound samples, a capability that is becoming increasingly accessible and affordable.
Imagine a scenario where, within hours of a wound assessment, clinicians can identify the precise E. coli strains present, their antibiotic resistance profiles, and their virulence potential. This information would allow for the selection of the most effective antibiotics, potentially combined with targeted therapies to disrupt biofilms or enhance the immune response. This isn’t science fiction; it’s a rapidly approaching reality.
Predictive Diagnostics and the Rise of AI
Beyond personalized treatment, the genomic data also opens the door to predictive diagnostics. By analyzing the genetic makeup of E. coli strains, researchers may be able to identify biomarkers – genetic signatures – that predict the likelihood of treatment failure or the risk of amputation. This would allow for proactive intervention, potentially preventing minor infections from escalating into life-threatening complications.
Artificial intelligence (AI) and machine learning are poised to play a crucial role in this area. AI algorithms can analyze vast datasets of genomic information to identify patterns and predict outcomes with greater accuracy than traditional methods. We can anticipate the development of AI-powered diagnostic tools that can assess DFI risk in real-time, guiding clinical decision-making.
| Metric | Current Status (2024) | Projected Status (2030) |
|---|---|---|
| Genomic Sequencing Time | 24-72 hours | Under 4 hours |
| Cost per Genome Sequence | $200 - $500 | $50 - $100 |
| AI-Driven Diagnostic Accuracy | 70% | 95% |
The Role of Phage Therapy and Novel Antimicrobials
As antibiotic resistance continues to rise, alternative therapies are gaining traction. Phage therapy – using viruses that specifically target and kill bacteria – is showing promising results in treating antibiotic-resistant infections. The genomic insights gained from studies like this one are crucial for identifying the most effective phages for specific E. coli strains. Furthermore, research into novel antimicrobials, including those that disrupt bacterial biofilms or target unique bacterial pathways, is accelerating.
Frequently Asked Questions About E. coli and Diabetic Foot Infections
What can I do to prevent E. coli infections in my feet if I have diabetes?
Meticulous foot care is paramount. This includes daily inspection for cuts or sores, washing your feet with mild soap and water, keeping your feet dry, and wearing properly fitting shoes. Promptly address any wounds, even minor ones, with appropriate wound care.
How will genomic sequencing impact my treatment if I develop a DFI?
Genomic sequencing will allow your doctor to identify the specific E. coli strains causing your infection and their antibiotic resistance profile. This will enable them to choose the most effective antibiotics, minimizing treatment failure and improving your chances of recovery.
Is phage therapy widely available for DFIs?
Phage therapy is still considered an experimental treatment in many regions, but it is gaining increasing attention and is available through specialized centers and clinical trials. Its availability is expected to expand as research progresses and regulatory hurdles are overcome.
The genomic revolution in understanding E. coli and diabetic foot infections is not just a scientific advancement; it’s a paradigm shift in patient care. By embracing personalized medicine, predictive diagnostics, and innovative therapies, we can significantly reduce the burden of this debilitating condition and improve the lives of millions living with diabetes. The future of DFI treatment isn’t about fighting bacteria with brute force; it’s about understanding them at a fundamental level and tailoring our approach accordingly.
What are your predictions for the role of AI in combating antibiotic resistance in DFIs? Share your insights in the comments below!
Discover more from Archyworldys
Subscribe to get the latest posts sent to your email.