For decades, neuroscience has been obsessed with the “wires”—the neurons and their synapses. But a new wave of high-resolution mapping is shifting the focus toward the brain’s “dark matter”: the astrocyte network. Recent methodological breakthroughs in volumetric imaging and genetic manipulation are finally allowing us to treat the brain not as a collection of slices, but as a complex, integrated circuit board where non-neuronal cells manage the flow of information and resources.
- Volumetric Precision: The integration of tissue clearing and light-sheet microscopy now allows for the mapping of astrocyte networks across whole-brain volumes, registered to the Allen Reference Atlas.
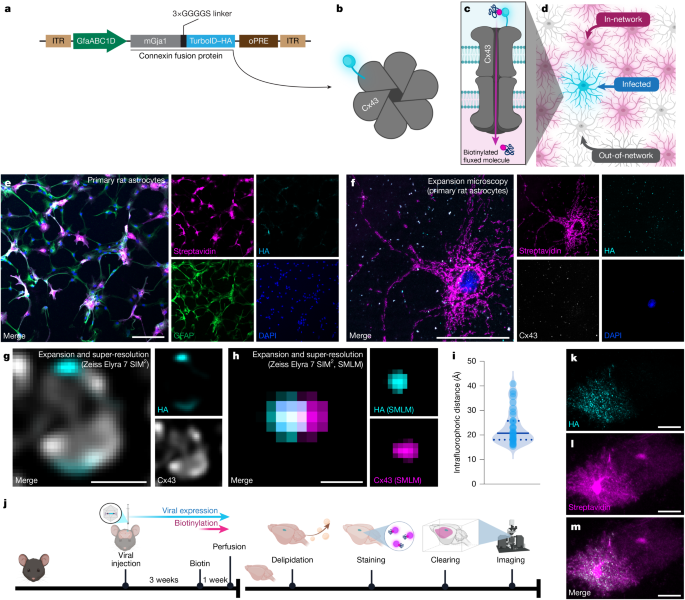
- Breaking the Resolution Barrier: By utilizing Expansion Microscopy (ExM) and Lattice-SIM, researchers are bypassing the diffraction limit of light to visualize gap junction proteins (Cx43 and Cx30) at a near-molecular scale.
- Dynamic Genetic Control: The use of tamoxifen-sensitive cre-ERT2 systems enables the “switching off” of specific proteins in adult mice, allowing scientists to observe the immediate systemic failure of cellular networks in real-time.
The Deep Dive: Reverse Engineering the Glial Grid
To understand why this methodology matters, you have to look at the “specs” of the imaging pipeline. Traditionally, studying the brain meant slicing it into thin sections—essentially trying to understand a 3D processor by looking at 2D photographs of its layers. The approach detailed here employs a brutal but effective sequence: delipidation (stripping the brain of fats), chemical clearing (making the tissue transparent), and light-sheet imaging.
The data overhead is staggering. A single reconstructed brain volume can average 280GB in size. To make sense of this “data swamp,” the team employs random-forest pixel classifiers and non-linear registration to the Allen Mouse Common Coordinate Framework (CCFv3). This isn’t just biology; it’s a massive data engineering project. They are mapping the “ports” of the astrocyte network—specifically the Gja1 and Gjb6 genes—to see how these cells communicate via gap junctions. By deleting these proteins, they can effectively “unplug” the glial network and measure the resulting cognitive and physiological fallout.
The Forward Look: Beyond the Neuron
The industry has spent billions trying to interface with neurons via electrodes (think Neuralink), but this research suggests we may be targeting the wrong hardware. If the astrocyte network is the primary regulator of synaptic plasticity and metabolic resource distribution, the next generation of Brain-Computer Interfaces (BCIs) might need to target glia, not just axons.
Watch for a pivot toward “Glial Therapeutics.” If we can map exactly how these networks fail in neurodegenerative diseases, we stop treating the symptoms (neuron death) and start treating the infrastructure (network collapse). The logical next step is the application of these 3D mapping techniques to human pathology samples, which will likely reveal that many “neuronal” diseases are actually “network” diseases. Expect a surge in “Connectomics 2.0” software—tools specifically designed to handle the terabyte-scale volumetric maps required to navigate the brain’s non-neuronal architecture.
Discover more from Archyworldys
Subscribe to get the latest posts sent to your email.