The cellular skeleton is no longer a static blueprint. For years, we’ve viewed F-actin—the primary structural filament of the cell—as a relatively predictable cable. But new data reveals a far more chaotic and dynamic reality: under mechanical stress, these filaments don’t just bend; they “supercoil,” twisting into complex geometries that likely dictate how cells move, adhere, and survive.
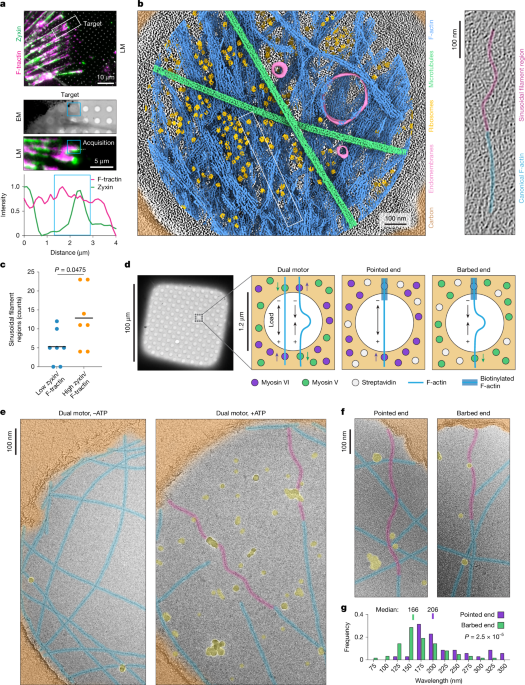
- The “Supercoil” Discovery: Researchers used dual-motor myosin systems to prove that compressive force triggers a specific, oscillatory “supercoiling” of actin filaments.
- AI as the Microscope: The study relied on Denoising Autoencoders (DAEs) and neural network particle pickers to extract near-atomic signals from the notoriously “noisy” environment of cryo-electron tomography (cryo-ET).
- Force-Activated Architecture: The data maps how $alpha$-catenin—a critical protein for cell-to-cell adhesion—reconfigures itself when the actin skeleton is under tension.
The Deep Dive: Breaking the “Noise” Barrier
To the uninitiated, this looks like a chemistry experiment. To a tech analyst, this is a masterclass in signal processing.
The fundamental problem with Cryo-ET is the signal-to-noise ratio. You are imaging biological matter at cryogenic temperatures, where the “noise” often drowns out the “signal” of the protein. The brilliance of this approach isn’t just in the hardware (the Titan Krios TEM), but in the computational layer.
The team implemented a Denoising Autoencoder (DAE) with adversarial training. Essentially, they taught an AI what a “perfect” actin filament looks like using synthetic data and then forced it to find those patterns in the messy, real-world experimental data. By using a “domain classifier” to bridge the gap between synthetic and real images, they effectively created a digital lens that can “see” through the ice and carbon contamination that usually ruins these samples.
Furthermore, the use of a “dual motor” condition—employing both Myosin V and Myosin VI—is a clever piece of biological engineering. By pitting motors that move in opposite directions against each other, the researchers created a controlled “tug-of-war,” allowing them to simulate the mechanical stresses a cell feels during migration or contraction in a vacuum of variables.
The Forward Look: Toward a “Mechanical Map” of the Cell
We are exiting the era of “static” structural biology and entering the era of mechanobiology.
The ability to map the $alpha$-catenin–F-actin complex under force is a precursor to something much larger. If we can determine exactly how the cell’s “glue” (adhesion proteins) reacts to mechanical torque, we can begin to model cancer metastasis with terrifying precision. Cancer cells are, at their core, masters of mechanical manipulation; they break their anchors and reshape their skeletons to squeeze through tissues.
What to watch for next:
- Whole-Cell Simulation: The “coarse-grained molecular dynamics” used here are a stepping stone. Expect a shift toward real-time, AI-driven simulations of entire cellular networks rather than single filaments.
- AI-Driven Drug Discovery: Once we have the “force-activated” structure of proteins like $alpha$-catenin, pharmaceutical companies will target these transient states—developing drugs that stabilize or destabilize cell adhesion based on mechanical triggers.
- The Hardware Race: As the AI processing catches up, the demand for higher-throughput Cryo-ET will skyrocket, likely leading to a new generation of “automated” microscopes that can identify and image “supercoil” events in real-time without human intervention.
Bottom line: The “hype” around AI in science is usually about LLMs writing papers. The actual value is here—in the trenches of data denoising and the mapping of the invisible forces that keep us held together.
Discover more from Archyworldys
Subscribe to get the latest posts sent to your email.