A new study detailing sophisticated methods for modeling and analyzing brain tumor development in mice – and, crucially, applying those methods to human samples – offers a significant leap forward in our understanding of these devastating cancers. The research, published in Nature, doesn’t present a single breakthrough cure, but rather a powerful toolkit for dissecting the complex interplay of genetics, cellular states, and tumor evolution. This is particularly important given the historically limited success rates in treating aggressive brain cancers like ependymoma.
- Precision Modeling: Researchers used in utero electroporation in mice to introduce specific genetic alterations linked to tumor development, allowing for controlled study of cancer initiation and progression.
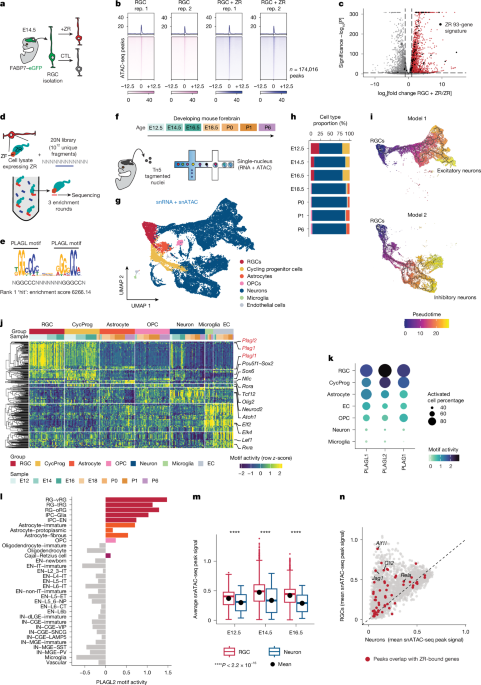
- Single-Cell Resolution: The study leverages cutting-edge single-nucleus multi-omics (snMultiome) technology – simultaneously analyzing RNA and chromatin accessibility – to map the cellular landscape of both mouse models and human tumors with unprecedented detail.
- Lineage Tracking: A novel “TrackerSeq” system allows researchers to trace the origins and evolutionary paths of cancer cells, revealing how they diverge from normal brain cells.
The Deep Dive: A Multi-Omics Approach to Brain Tumor Complexity
Brain tumors, particularly those arising in children, are notoriously difficult to treat. Their heterogeneity – the fact that they are composed of diverse cell types and genetic mutations – contributes to treatment resistance and relapse. Traditional methods often struggle to capture this complexity. This study addresses this challenge head-on by combining several advanced techniques. In utero electroporation allows for precise genetic manipulation in a developing brain, creating a controlled environment to study tumor formation. The use of snMultiome technology is a game-changer. By analyzing the RNA and chromatin accessibility of individual nuclei, researchers can identify the specific genes and regulatory elements that are active in different cell types within the tumor. This provides a far more nuanced picture than bulk tissue analysis.
The development and application of TrackerSeq is particularly noteworthy. This system allows researchers to “barcode” cells, enabling them to trace their lineage – essentially, to reconstruct the family tree of the tumor. This is critical for understanding how cancer cells evolve and adapt, and for identifying the root causes of tumor growth. The researchers focused on the role of the ZFTA–RELA fusion gene, a common driver of aggressive ependymoma, and the PLAGL1 gene, implicated in glioma development. Their work reveals how these genes disrupt normal brain development and drive the formation of malignant cells.
The Forward Look: Towards Targeted Therapies and Early Detection
While this study is primarily a foundational effort, its implications for future cancer treatment are significant. The detailed cellular maps and lineage tracing data generated by this research will serve as a valuable resource for identifying new drug targets. Specifically, understanding the disrupted regulatory networks controlled by ZFTA–RELA and PLAGL1 could lead to the development of therapies that specifically block their oncogenic activity. The identification of progenitor and lineage scores for malignant cells offers a potential biomarker for assessing tumor aggressiveness and predicting treatment response.
However, the most immediate impact may be in refining existing therapeutic strategies. The ability to identify and characterize different subtypes of brain tumors, based on their molecular signatures and lineage, could allow for more personalized treatment approaches. Furthermore, the insights gained from this study could inform the development of early detection methods. If we can identify the earliest signs of tumor development – perhaps by detecting changes in gene expression or chromatin accessibility – we may be able to intervene before the tumor becomes aggressive and difficult to treat. The researchers’ reporting summary highlights the rigor of their methodology, bolstering confidence in the reproducibility and reliability of these findings. Expect to see this toolkit adopted by other research groups, accelerating the pace of discovery in the field of brain cancer research. The next phase will likely involve translating these findings into clinical trials, testing the efficacy of new therapies in patients.
Discover more from Archyworldys
Subscribe to get the latest posts sent to your email.