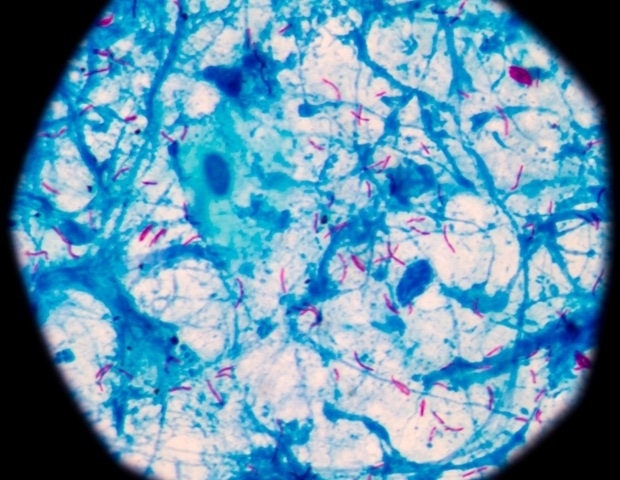
The global fight against tuberculosis (TB) has received a significant boost with new research detailing how a promising class of experimental antibiotics disrupts the bacterium responsible for the disease. This isn’t simply a discovery of *what* these antibiotics do, but *how* – a crucial distinction that accelerates the development of desperately needed new treatments, particularly against drug-resistant strains.
- Novel Mechanism Uncovered: Researchers pinpointed how ecumicin, ilamycin, and cyclomarin interfere with the ClpC1–ClpP1P2 protein degradation complex in Mycobacterium tuberculosis.
- Beyond Simple Shutdown: The antibiotics don’t just halt the bacterial recycling system; they create imbalances that weaken the bacterium’s ability to survive.
- Expanded Treatment Pipeline: This research offers a pathway to refine existing compounds and design more effective anti-TB drugs, including options for drug-resistant TB.
TB remains a devastating global health crisis, claiming 1.2 million lives annually. The emergence of drug-resistant TB is particularly alarming, rendering existing treatments ineffective and fueling the spread of the disease, especially in regions like the Asia-Pacific. The urgency stems from a decades-long stagnation in TB drug development. The last major anti-TB drug, bedaquiline, was approved in 2012, highlighting the critical need for innovation.
The study, published in Nature Communications, focuses on the ClpC1–ClpP1P2 complex – essentially the bacterium’s internal recycling system. This complex breaks down damaged proteins, allowing the TB bacterium to adapt and survive under stress, such as within the human body. Targeting this system is attractive because its disruption is lethal to the bacterium. What’s particularly insightful about this research is the nuanced understanding of *how* the three antibiotics interact with this complex. Previous research often focused on simply identifying compounds that inhibited the complex, but this study reveals each antibiotic has a unique disruptive effect, creating widespread internal imbalances.
Researchers meticulously tracked changes in over 3000 proteins within the bacterium, providing an unprecedented view of the ripple effects caused by disrupting the ClpC1–ClpP1P2 complex. This comprehensive approach, as explained by first author Isabel Barter, allows for a more strategic refinement of these compounds.
The Forward Look
The identification of the ClpC1–ClpP1P2 complex as a promising drug target is a pivotal step, but several key developments are now anticipated. First, we can expect intensified research focused on optimizing these three compounds – ecumicin, ilamycin, and cyclomarin – to maximize their disruptive effects. This will likely involve structural modifications to enhance their binding affinity to the ClpC1–ClpP1P2 complex and improve their delivery to the site of infection. Second, pharmaceutical companies will likely increase investment in this area, spurred by the detailed mechanistic understanding provided by this study. Clinical trials, initially focusing on safety and efficacy in small patient cohorts, are likely to begin within the next 2-3 years. Finally, the research team is already exploring other compounds that might target this complex, potentially leading to a broader range of anti-TB drug candidates. The success of this approach could not only revitalize TB treatment but also provide a blueprint for tackling other bacterial infections by targeting similar protein degradation systems.
Discover more from Archyworldys
Subscribe to get the latest posts sent to your email.